 Zhijun Wang
Zhijun Wang
Associate professor
DNA phosphorothioation
Email: wangzhijun@sjtu.edu.cn
DNA Phosphorothioate Modification as an Antioxidant
Endogenous superoxide and hydrogen peroxide are generated as by-products of aerobic metabolism. The molecular damages accumulated from these reactive oxygen species is one of the causes leading to aging and diseases. Reactive oxygen species scavenging enzymes and small molecules have been being evolved to protect cells.
A newly finding is, diverse bacteria contain DNA with sulfur incorporated stereo-specifically into their DNA backbone, called phosphorothioate modification. Phosphorothioate DNA can react with hydrogen peroxide or peracetic acid and the reaction has two possible outcomes: DNA backbone cleavage or sulfur removal resulting in restoration of normal DNA backbone. The phosphorothioate DNA hosting bacteria show increasing resistance to hydrogen peroxide. Inside the cells, phosphorothioate DNA is more resistant to the double-strand break damage caused by hydrogen peroxide than phosphorothioate-free DNA. These findings are consistent with a hypothesis that phosphorothioation modification endows DNA with reducing chemical property, protecting the bacteria against reactive oxygen species.
Education Experience
2002-2007 Ph.D. in Biochemistry and Molecular Biology,
The Institute of Microbiology, Chinese Academy of Sciences,
With Prof. Keqian Yang.
1994-1997 M.S. in Molecular Plant Biochemistry,
Nanjing Agricultural University,
With Prof. Maobing Ye.
1986-1990 B.S. in Plant Protection Science,
Nanjing Agricultural University
Work Experience
2011.06- Tutor for graduate, SJTU, China.
2011.02-2013.02 Tang Berkeley Scholar,
The Department of Molecular & Cell Biology (MCB),
University of California, Berkeley.
2010.12- Associate Professor, SJTU, China.
2007.01-2010.12 Assistant professor, SJTU, China.
1997.06-2002.09 Lecturer, Department of Chemical Engineering,
Nantong University.
Research Interests
1. Anti-oxidation function of bacteria DNA phosphorothioate modification and its application.
2. The structure and function of calcimycin biosynthetic enzyme complex.
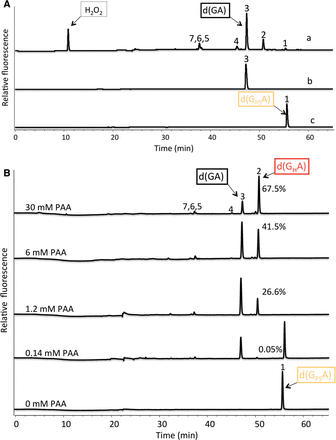
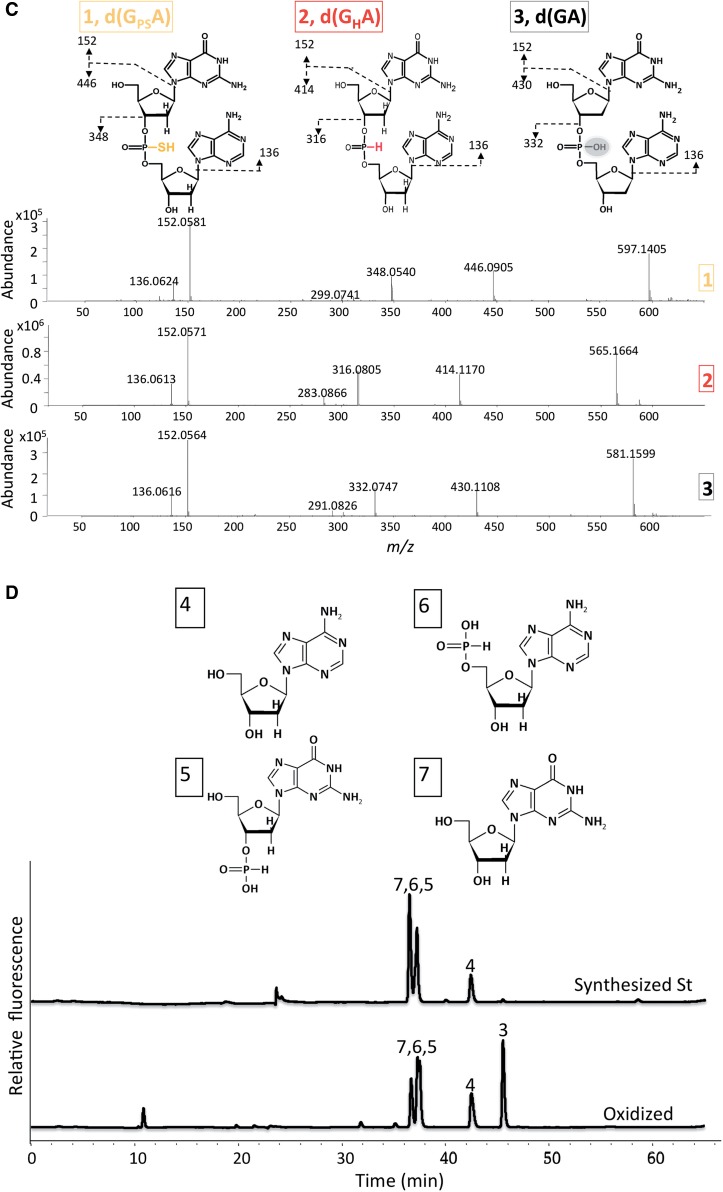
Figure 1 PT dinucleotide is oxidized by H2O2 and PAA. (Xie , et al. Nucleic Acids Res. 2012)
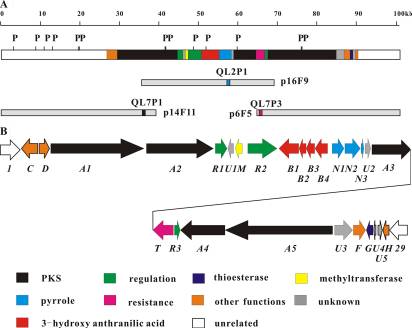
Figure 2 Calcimycin biosynthesis gene cluster from S. chartreusis NRRL 3882.(Wu et al. Antimicrob Agents Chemother. 2011)
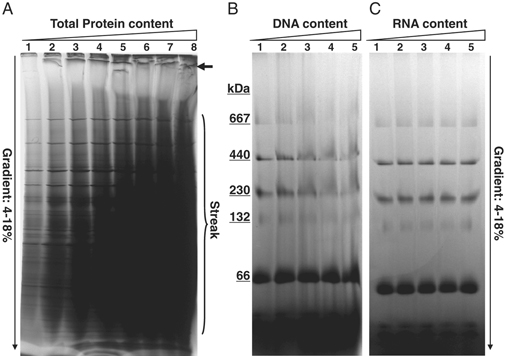
Figure 3 Interference by DNA in protein samples in BN-PAGE. (Liang, et al. Electrophoresis. 2009 )
Selected Publications
1. Characterization of the N-methyltransferase CalM involved in calcimycin biosynthesis by Streptomyces chartreusis NRRL 3882. Wu QL, Gou LX, Lin SJ, Liang JD, Yin J, Zhou XF, Bai LQ, An DR, Deng ZX*, Wang ZJ*. Biochimie. 2013 Apr 11.
2. Mutasynthesis of Pyrrole Spiroketal Compound by Using Calcimycin 3-Hydroxy Anthranilic Acid Biosynthetic Mutant. Gou LX, Wu QL, Lin SJ, Li XM, Liang JD, Zhou XF, An DR*, Deng ZX*, Wang ZJ*. Applied Microbiology and Biotechnology ,2013 May 12.
3. Phosphorothioate DNA as an antioxidant in bacteria. Xie XQ, Liang JD, Pu TN, Xu F, Yao F, Yang Y, Zhao YL, You DL, Zhou XF, Deng ZX*, Wang ZJ*. Nucleic Acids Research. 2012 Oct; 40(18):9115-24.
4. Characterization of the Pyrrole Polyether Antibiotic Calcimycin (A23187) Biosynthetic Gene Cluster from Streptomyces chartreusis NRRL 3882. Wu QL, Liang JD, Lin SJ, Zhou XF, Bai LQ *, Deng ZX and Wang ZJ*. Antimicrobial Agents and Chemotherapy (AAC),2011 Mar; 55(3):974-82.
5. Effective elimination of nucleic acids from bacterial protein samples for optimized blue native polyacrylamide gel electrophoresis. Liang JD, Niu QL, Xu XP, Luo YM, Zhou XF, Deng ZX*, Wang ZJ*. Electrophoresis.2009 Jul; 30(14):2454-9.
6. Localization of the ActIII actinorhodin polyketide ketoreductase to the cell wall. Xu XP#,Wang ZJ#,Fan KQ,Wang SL,Jia CJ,Han H,Ramalingam E,Yang KQ. FEMS MicrobiolLett.2008 Oct; 287(1):15-21.
7. DNA modification by sulfur: analysis of the sequence recognition specificity surrounding the modification sites. Liang JD#, Wang ZJ#, He XY, Li JL, Zhou XF,Deng ZX. Nucleic Acids Research.2007 Apr; 35(9):2944-54.
8. Sample preparation for two-dimensional blue native/SDS polyacrylamide gel electrophoresis in the identification of Streptomyces coelicolor cytoplasmic proteincomplexes. Wang ZJ, Xu XP, Fan KQ, Jia CJ, Yang KQ. J Biochem Biophys Methods.2007 Jun 10; 70(4):565-72.
9. A novel target of IscS in Escherichia coli: participating in DNA phosphorothioation. An XH, Xiong W, Yang Y, Li FH, Zhou XF, Wang ZJ, Deng ZX, Liang JD. PLoS One. 2012; 7(12):e51265.
10. Theoretical study on steric effects of DNA phosphorothioation: B-helical destabilization in Rp-phosphorothioated DNA . Zhang Y, Liang J, Lian P, Han Y, Chen Y, Bai L, Wang Z, Liang J, Deng Z, Zhao YL. J Phys Chem B.2012 Sep 6;116(35):10639-48.
11. Crystal structure of the cysteine desulfurase DndA from Streptomyces lividans which is involved in DNA phosphorothioation. Chen F, Zhang Z, Lin K, Qian T, Zhang Y, You D, He X, Wang Z, Liang J, Deng Z, Wu G. PLoS One. 2012; 7(5):e36635.
12. Structural insights into DndE from Escherichia coli B7A involved in DNA phosphorothioation modification. Hu W, Wang C, Liang J, Zhang T, Hu Z, Wang Z, Lan W, Li F, Wu H, Ding J, Wu G, Deng Z, Cao C. Cell Res.2012 Jul;22(7):1203-6.
13. Purification, crystallization and preliminary X-ray analysis of the DndE protein from Salmonella enterica serovar Cerro 87, which is involved in DNA phosphorothioation. F. Chen, K. Lin, Z. Zhang, L. Chen, X. Shi, C. Cao, Z. Wang, J. Liang, Z. Deng, G. Wu. Acta Crystallogr Sect F Struct Biol Cryst Commun.2011 Nov 1; 67(Pt 11):1440-2.
14. Oxazolomycin biosynthesis in Streptomyces albus JA3453 featuring an "acyltransferase-less" type Ipolyketide synthase that incorporates two distinct extender units. Zhao C, Coughlin JM, Ju J, Zhu D, Wendt-Pienkowski E, Zhou X, Wang Z, Shen B, Deng Z. J Biol Chem.2010 Jun 25;285(26):20097-108
15. Two pHZ1358-derivative Vectors for Efficient Gene Knockout in Streptomyces. He, Y, Wang Z, Bai L, Liang J, Zhou X, Deng Z. J. Microbiol. Biotechnol. 2010 4; 20(4): 678-682
16. DNA phosphorothioation in Streptomyces lividans: Mutational analysis of the dnd locus. Xu T, Liang J, Chen S, Wang L, He X, You D, Wang Z, Li A, Xu Z, Zhou X, Deng Z. BMC Microbiol.2009 Feb 20; 9(1):41.
17. Characterization of a hyperthermostable Fe-superoxide dismutase from hot spring. He Y, Fan K, Jia C, Wang Z, Pan W, Huang L, Yang K,Dong Z. Appl Microbiol Biotechnol. 2007 75, 367-376.
Honors
Tang Berkeley Scholar
http://qb3.berkeley.edu/qb3/tang-wzj.cfm
Copyright © 2016,The Laboratory of Molecular Microbiology at Shanghai Jiao Tong University. All rights reserved.
Add:Science Building, Xuhui Campus, Shanghai Jiaotong University, Shanghai 200030, China. Tel:021-62932943
