 Hong-Yu Ou
Hong-Yu Ou
Professor
Microbial Genomics and Bioinformatics
Email: hyou@sjtu.edu.cn
The website of the Microbial Genomics and Bioinformatics Group available at http://bioinfo-mml.sjtu.edu.cn/.
Hong-Yu OU received his B.S. in Food Engineering at Nanjing Agricultural University in 1997, M.S. in Fermentation Engineering at Tianjin University of Science & Technology in 2001, and Ph.D. in Bioinformatics at Tianjin University in 2004. After finished a two-year postdoc at the University of Leicester, he began his academic career as an Associate Professor at Shanghai Jiao Tong University. In 2012, he was appointed as a Professor of Microbiology at the State Key Laboratory of Microbial Metabolism, Shanghai Jiao Tong University. His group’s research focuses on bacterial comparative genomics and bioinformatics. His current work emphasizes characterization of mobile genetic elements in Klebsiella pneumoniae and other opportunistic bacterial pathogens that have developed antibiotic resistance, and identification of antibiotic biosynthetic gene clusters with genome mining in Streptomyces. Over the years he has developed several bioinformatics tools and databases of bacterial mobile genetic elements, such as the mGenomeSubtractor tool for in silico subtractive hybridization, MobilomeFINDER for genomic island detection, the ICEberg database of integrative and conjugative elements, TADB of type 2 toxin-antitoxin loci, SecReT4 of type IV secretion system.
Research Interests:
1. Genome mining of Klebsiella pneumoniae and other opportunistic pathogens that have developed antibiotic resistance,
2. Identification of antibiotic biosynthetic gene clusters of Streptomyces,
3. Development and maintenance of bioinformatics tools and databases focused on bacterial mobile genetic elements.
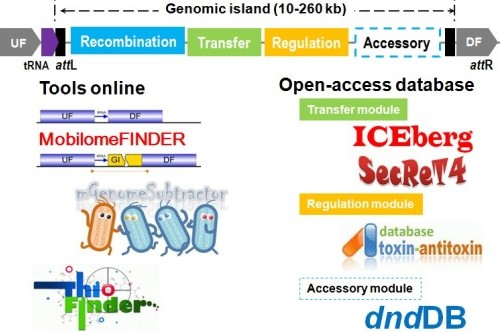
Figure 1. Bioinformatics tools and databased available at the Microbial Genomics and Bioinformatics Group, MML, SJTU.
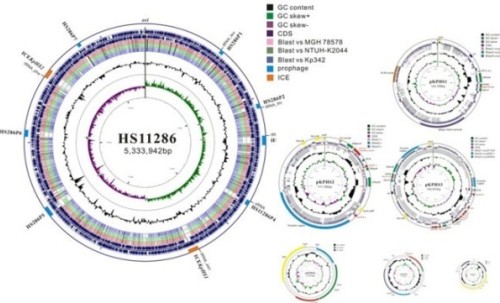
Figure 2. Complete genome sequence of Klebsiella pneumoniae subsp. pneumoniae HS11286, a multidrug-resistant strain belonging to the dominant clone ST11 of KPC-producing Klebsiella pneumoniae in China. [Liu, et al. Journal of Bacteriology, 2012]

Figure 3. Streptomyces cattleya is unusual for its capacity to biosynthesis of antibiotics and other secondary metabolites, such as fluoroacetate and epsilon-poly-L-Lysine biosynthesis. The mega-plasmid pSCATT is densely packed with 12 putative secondary metabolite gene clusters, in addition to the chromosome that also harbors 17 such gene clusters. [Zhao, et al. Bioorganic Chemistry, 2012]
Selected Publications:
1. SecReT4: a web-based bacterial type IV secretion system resource. Bi D, Liu L, Tai C, Deng Z, Rajakumar K*, Ou HY*, Nucleic Acids Research, 2013, 41: D660-D665.
2. Book chapter: “Type II Loci: Phylogeny”. Ou HY*, Wei Y, Bi D, Prokaryotic Toxin-Antitoxins, Editor Kenn Gerdes, 2013, Springer.
3. Bioinformatics tools and databases focused on genomic islands and secondary metabolite biosynthesis of Streptomyces. Ou HY*, Microbiology China, 2013, Actinomycetes issue (in Chinese).
4. ICEberg: a web-based resource for integrative and conjugative elements found in Bacteria. Bi D, Xu Z, Harrison E, Tai C, Wei Y, He X, Jia S, Deng Z, Rajakumar K*, Ou HY*, Nucleic Acids Research, 2012, 40: D621-D626.
5. Complete Genome Sequence of Klebsiella pneumoniae subsp. pneumoniae HS11286, a Multidrug-Resistant Strain Isolated from Human Sputum. Liu P, Li P, Jiang X, Bi D, Xie Y, Tai C, Deng Z, Rajakumar K, H.Y. Ou*, Journal of Bacteriology, 2012, 194: 1841-1842.
6. ThioFinder: a web-based tool for the identification of thiopeptide gene clusters in DNA sequences. Li J, Qu X, He X, Duan L, Wu G, Bi D, Deng Z, Liu W*, Ou HY*, PloS ONE, 2012, 7: e45878.
7. Insights into fluorometabolite biosynthesis in Streptomyces cattleya DSM46488 through knockout mutants. Zhao C, Li P, Deng Z, Ou HY*, McGlinchey RP, O'Hagan D*, Bioorganic Chemistry, 2012, 44: 1-7.
8. Book chapter: “Identification of bacterial genomic islands by comparative genomic analysis. Ou HY*, Microbiology genomics in China, Editor Zi-Niu Yu, 2011, Science Press (in Chinese).
9. TADB: a web-based resource for Type 2 toxin-antitoxin loci in bacteria and archaea. Shao Y, Harrison EM, Bi D, Tai C, He X, Ou HY*, Rajakumar K, Deng Z, Nucleic Acids Research, 2011, 39: D606-D611.
10. Expansion of the known Klebsiella pneumoniae species gene pool by characterization of novel alien DNA islands integrated into tmRNA gene sites. Zhang J, Jurriaan van Aartsen J, Jiang X, Shao Y, Tai C, He X, Tan Z, Deng Z, Jia S*, Rajakumar K, H.Y. Ou*, Journal of Microbiological Methods, 2011, 84: 283-289.
11. Book chapter: “ArrayOme- & tRNAcc-facilitated mobilome discovery: comparative genomics approaches for identifying rich veins of bacterial novel DNA sequences”. Ou HY, Rajakumar K, Handbook of Molecular Microbial Ecology I: Metagenomics and Complementary Approaches, Editor F. J. de Bruijn, 2011, John Wiley & Sons, Inc.
12. Cleavage of Phosphothioated DNA and Methylated DNA by the Type IV Restriction Endonuclease ScoMcrA. Liu G, Ou HY, Wang T, Li L, Tan H, Zhou X, Rajakumar K, Deng Z*, He X*, PLoS Genetics, 2010, 6: e1001253.
13. mGenomeSubtractor: a web-based tool for parallel in silico subtractive hybridization analysis of multiple bacterial genomes. Shao Y, He X, Tai C, Ou HY*, Rajakumar K, Deng Z, Nucleic Acids Research, 2010, 38: W194-W200.
14. The pheV phenylalanine tRNA gene in Klebsiella pneumoniae clinical isolates is an integration hotspot for possible niche-adaptation genomic islands. Chen N¶, Ou HY¶, van Aartsen JJ, Jiang X, Li M, Yang Z, Wei Q, Chen X, He X, Deng Z, Rajakumar K, Lu Y*, Current Microbiology, 2010, 60, 210-6. (¶ These authors contributed equally to this work.)
15. dndDB: a database focused on phosphorothioation of the DNA backbone. Ou HY, He X, Shao Y, Tai C, Rajakumar K, Deng Z*, PLoS ONE, 2009, 4:e5132.
16. Translational Genomics to Develop a Salmonella enterica Serovar Paratyphi A Multiplex PCR Assay. Ou HY¶, Ju CTS ¶, Thong KL, Ahmad N, Deng Z, Barer MR, Rajakumar K*, Journal of Molecular Diagnostics, 2007, 9: 624-630. (¶ These authors contributed equally to this work.)
17. MobilomeFINDER: web-based tools for in silico and experimental discovery of bacterial genomic islands. Ou HY, He X, Harrison EM, Kulasekara BR, Thani AB, Kadioglu A, Lory S, Hinton JC, Barer MR, Deng Z*, Rajakumar K*, Nucleic Acids Research, 2007, 35, W97-W104.
18. Analysis of a genomic island housing genes for DNA S-modification system in Streptomyces lividans 66 and its counterparts in other distantly related bacteria. He X, Ou HY, Yu Q, Zhou X, Wu J, Liang J, Zhang W, Rajakumar K, Deng Z.*, Molecular Microbiology, 2007, 65, 1034-48.
19. A novel strategy for identification of genomic islands by comparative analysis of the contents and contexts of tRNA sites in closely related bacteria. Ou HY, Chen LL, Lonnen J, Chaudhuri RR, Thani AB, Smith R, Garton NJ, Hinton JC, Pallen M, Barer M, Rajakumar K*, Nucleic Acids Research, 2006, 34: e3.
20. ArrayOme: a program for estimating the sizes of the microarray-visualised genomes. Ou HY, Smith R, Lucchini S, Hinton JC, Chaudhuri RR, Pallen M, Barer M, Rajakumar K*, Nucleic Acids Research, 2005, 33: e3.
Copyright © 2016,The Laboratory of Molecular Microbiology at Shanghai Jiao Tong University. All rights reserved.
Add:Science Building, Xuhui Campus, Shanghai Jiaotong University, Shanghai 200030, China. Tel:021-62932943
